Spectroscopic studies of lichen depsides and depsidones
Jose Vila1, Patricia Mollinedo1, Olov Sterner2
1Instituto de Investigaciones en Productos Naturales, Carrera de Ciencias Químicas, 2º piso Edificio Facultativo FCPN, calle 27 y calle Andrés Bello, Campus Universitario Cota Cota, Universidad Mayor de San Andrés, La Paz-Bolivia.
2Division of Organic Chemistry, Lund University, PO Box 124, 221 00 Lund Sweden
Keywords: Lichen, depside, depsidone, structure determination
ABSTRACT
A structural study of depsides and depsidones by 1H NMR and 13C NMR spectroscopy, in which assignments of all carbons/protons of compounds isolated from various lichens (Thamnolia vermicularis, Baeomyces roseus and Everniastrum cirrhatum) is reported. The compounds studied include thamnolic acid, decarboxythamnolic acid, baeomycesic acid, salazinic acid and atranorin. The assignments are based on variuos 1D and 2D NMR experiments, e.g. DEPT, COSY, HMQC and HMBC.
Corresponding author: pattymollinedo@gmail.com
Lichens are complex organism consisting of a symbiotic association of an alga and a fungus.[1],[2],[3] The people in Bolivia have for centuries used herbal medicine for different purposes, one is based on the lichen Thamnolia vermicularis and used for the treatment of cough and hoarseness.[4] Depsides and depsidones are important groups of secondary metabolites of lichens that often are present in high concentrations, and besides their medicinal properties they are also believed to be important for the ability of the lichens to defend themselves to the oxidative stress caused by solar irradiation. As part of a general study of depsides and depsidones, we collected three different species of lichens, Thamnolia vermicularis, Baeomyces roseus and Everniastrum cirrhatum, from the area of Pongo 3800 m.a.s.l. (La Paz Bolivia). From Everniastrum cirrhatum, atranorin (1) was found to be the main component while salazinic acid (3) is a minor compound. Decarboxythamnolic acid (5) was isolated as a minor component and thamnolic acid (6) as the main component from Thamnolia vermicularis, while baeomycesic acid (7) was isolated as a main component from Baeomyesis roseus. A new derivate of atranorin was prepared and characterized by NMR spectroscopy. A previous NMR investigation reported assignments of 13C NMR of depsides, depsidones and depsones.[5],[6] However, as this was carried out without the use of two-dimensional NMR techniques some assignments of thamnolic acid, baeomycesic acid and hexaacetylsalazinic acid are incorrect. The correct shifts are presented in this investigation.
RESULTS, DISCUSSION
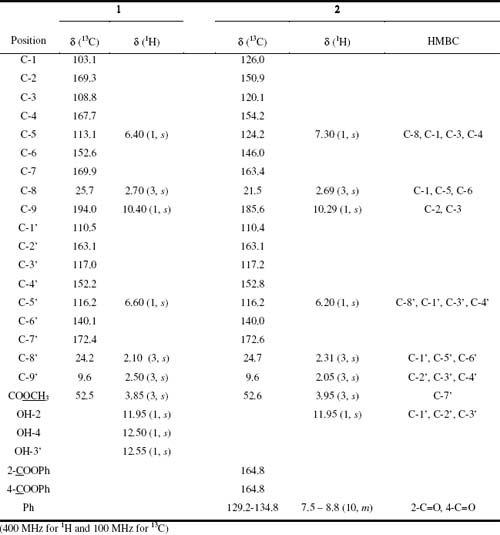
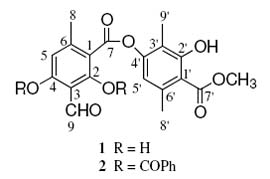
Atranorin (1) was isolated from Everniastrum cirrhatum, and was obtained as white crystals (m.p. 197-198 oC from CH2Cl2-EtOAc mixture). The molecular formula C19O8H18 was deduced from the analysis of FABMS and NMR data. The 1H NMR spectrum of 1 displayed signals for three methyl groups at d 2.11 s, d 2.59 s, d 2.71 s and one group methoxyl at d 4.05 s. Two aromatic protons at d 6.44 s and d 6.56 s and four signals at d 10.39, d 11.95, d 12.53 and d 12.59 were also observed. The analysis of the 13C NMR spectrum data of 1 with the aid of HMQC and HMBC spectrum permitted the complete structure assignment of para-depside as atranorin (Table 1). 2,4-bis(Benzoyloxy)-atranorin (2) was obtained by esterfication of 1 with benzoyl chloride in chloroform and in the presence of pyridine. The 1H NMR spectrum of 2 show signals of four methyl groups at d 2.05 s, d 2.31 s, d 2.69 s and oxygenated methyl at d 3.95 s, as well as two proton aromatic at d 6.20 s and d 7.30 s. Other signals in the aromatic region corresponding to the two phenyl groups; at d 7.55 m (4H), d 7.75 m (2H) and d 8.25 m (4H), an aldehyde proton at d 10.29 s and a hydrogen bonding OH proton at d 11.95 s. The 13C NMR spectrum indicated the presence of four methyl, two protonated unsaturated carbons and twelve non-protonated, as well as two carbonyl groups at 164.8 ppm, corresponding to a para-depside skeleton, in addition to the signals of the two phenyls. After the assignment of all protons to their respective carbons from HMQC experiments, a HMBC experiment was performed to complete the structural elucidation of 2. In this, cross correlation peaks were observed between H-5 (d 7.30, s, 1H) and C-8, C-1, C-3 and C-4; from H-8 (d 2.69, s, 3H) and C-1, C-5 and C-6; from H-8 (d 2.31, s, 3H) and C-1, C-5 and C-6; from H-9 (d 2.05, s, 3H) and C-2, C-3 and C-4; from H-5 (d 6.20, s, 1H) and C-8, C-1, C-3 and C-4; from the methoxyl protons (d 3.95, s, 3H) and C-7; and from the aldehyde proton H-9 (d 10.29, s, 1H) and C-2 and C-3. In addition, correlations were observed between the 2-OH proton (d 11.95, s, 1H) and C-1, C-2 and C-3. The HMBC correlations are summarized in Table 1. Thus, the structure of compound 2 was established as 2,4-bis(benzoyloxy)-atranorin.
Table 1. 13C and 1H NMR data for Atranorin 1 and 2,4-bis(benzoyloxy)-atranorin 2 and HMBC for 2

]]> The elemental composition of decarboxythamnolic acid (3) was suggested to be C18H16O9, from FABMS ([M+1] at m/z 377), 1H NMR and 13C NMR data. The 1H NMR spectrum (DMSO-d6) showed signals for three methyl groups with singlets at d 2.28 s, d 2.50 s and d 3.90 s, two aromatic protons at d 6.38 s, d 6.65 s, as well as an aldehyde proton at d 10.23 s. The 13C NMR and DEPT-135 spectra confirmed the presence of eighteen carbons, three methyls, three protonated unsaturated carbons and twelve non-protonated.
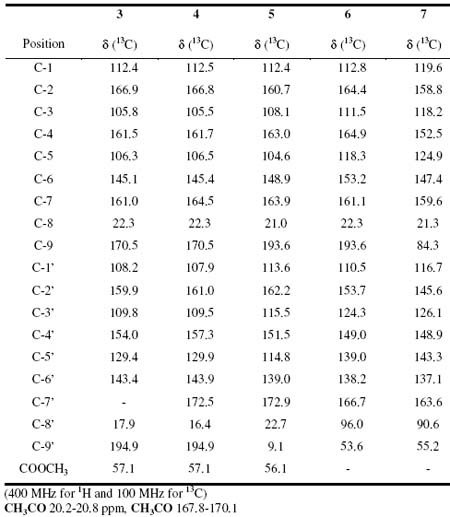
The complete structural elucidation of 3 was achieved by analysis of the HMBC. Pertinent HMBC correlations were observed between protons and carbons: from H-8 (d 2.50, s, 3H) with C-1 (d 112.4), C-5 (d 106.3) and C-6 (d 145.1), between H-5 (d 6.65, s, 1H) and C-1 (d 112.4), C-3 (d 105.8) and C-8 (d 22.3), between H-8 (d 2.28, s, 3H) and C-1 (d 108.2), C-5 (d 129.4) and C-6 (d 143.4), between H-1 (d 6.38, s, 1H) and C-8 (d 17.9), C-3 (d 109.8), C-5 (d 129.4) and C-2 (d 159.9), between H-9 (d 10.23, s, 1H) and C-4 (d 154.0) and betwwen CH3O (d 3.90, s, 3H) and C-4 (d 161.5). The structure was established on the basis of the above data as meta-depside decarboxythamnolic acid (Table 2). The elemental composition of thamnolic acid (4) was determined to be C19H16O11 from FABMS ([M+1] at m/z 421) and 1H NMR and 13C NMR. The 1H NMR (DMSO-d6) spectrum showed signals for three methyl groups at with singlets d 2.45 s, d 2.50 s and d 3.86 s, an aromatic proton at d ]]> s, and an aldehyde proton at d 10.4 s. The 13C NMR and DEPT-135 spectra confirmed the presence of nineteen carbon atoms three methyls, two protonated unsaturated carbons and fourteen non-protonated. In the HMBC spectrum correlations were observed between H-8 (d 2.50, s, 3H) and C-1 (d 112.5), C-5 (d 106.5) and C-6 (d 145.4), between H-5 (d 6.15, s, 1H) and C-1 (d 112.5), C-3 (d 105.5) and C-8 (d 22.3), between H-8 (d 2.45, s, 3H) and C-1 (d 16.4), C-5 (d 129.9) and C-6 (d 143.9). Further correlations were observed between H-9 (d 10.30, s, 1H) and C-4 (d 157.3), C-5 (d 129.9), and C-3 (d 109.5), and between CH3O (d 3.86, s, 3H) and C-4 (d 161.7). There were still four signals of carbons additional remaining in the 13C RMN spectrum at d 161.0, d 157.3, d 166.8 and d 164.5. ]]> 3, because decarboxythamnolic acid showed similar chemical shift of carbon atoms which the compound 3 and were assigned to: C-2 (d 161.0), C-4 (d 157.3), C-2 (d 166.8) and C-7 (d 164.5), 4 could be identified as meta-depside thamnolic acid (Table 2). Baeomycesic acid (5) was obtained as a solid. The elemental composition C19H18O8 was deduced from FABMS ([M+1] at m/z 375) and NMR data analysis. The 1H NMR (DMSO-d6) spectrum contained seven signals, three of them are methyl groups at d 2.02 s, d 2.47 s and d 2.51 s, and one methoxy group at d 3.96 s. Two aromatic protons at d 6.45 s, d 6.69 s, a proton from aldehyde ]]> s. The 13C NMR spectrum indicated the presence of nineteen carbons atoms. These observations were consistent with C19H18O8, the analysis of the 13C NMR spectrum revealed the presence of four methyls, three protonated unsaturated carbons and twelve non-protonated. The complete structural elucidation of 5 was achieved by analysis of the HMQC and HMBC. Important HMBC cross correlation peaks were observed between the following protons and carbons: d 2.49 (3H, s, H-8) and C-1 (d 112.4), C-5 (d 104.6) and C-6 (d 148.9); d 6.69 (1H, s, H-5) and C-1 (d 112.4) and C-8 (d 21.0); d 2.51 (3H, s, H-8) and C-1 (d 113.6), C-5 (d 114.8) and C-6 (d 139.0); d 2.05 (3H, s, H-9) and C-2 (d 162.2), C-3 (d 115.5) and C-4 (d 151.5); d 6.45 (1H, s, H-5) and C-1 (d 113.6), C-3 (d 115.5), C-4 (d 151.5) and C-8 (d 22.7); d 10.21 (1H, s, H-9) and C-2 (d 160.7) and C-3 (d 108.1); and CH3O d 3.96 (3H, s) and C-4 (d 163.0), were observed. The structure was established as the para-depside baeomycesic acid (Table 2).The FABMS of salazinic acid (6) showed a molecular ion at ([m+1] at m/z 389), while the 1H NMR and 13C NMR spectra indicated that the molecular formula should be C18H12O10. The 1H NMR (DMSO-d6) spectrum showed signals for one methyl group at d 2.51 s, one CH2OH group at d 4.58 s and two signals at d 6.81 s and d 6.78 s, as well as an aldehyde proton at d 10.23 s. ]]> 13C NMR and DEPT-135 experiment spectrum indicated the presence of eighteen carbons atoms, the compound had thirteen non-protonated unsaturated carbons, three protonated unsaturated carbons, one mono-protonated saturated carbon and one di-protonated saturated carbon, in addition to a methyl. The analysis of HMQC, COSY and HMBC spectra permitted the complete structure assignment of the depsidone salazinic acid (6), previously isolated from Parmotrema delicatulum[7] (Table 2). The acetylation of 6 with acetic anhydride in pyridine gave hexaacetylsalazinic acid (7) as the sole product. The elemental composition of 7 was suggested by FABMS ([M+1] m/z 659), the sodium adduct [M+Na] was also observed at m/z 681 and NMR data to be C30H26O17. The 1H NMR (CDCl3) spectrum showed signals between d 1.9 2.7 for seven methyl groups, one methylen group at d 5.1 (H-8a, d, J=7.0 Hz) and d 5.4 (H-8b, d, J=7.0 Hz), an aromatic proton at d 6.9 s and two methine protons at d 7.8 s and d 7.95 s. The 13C NMR and DEPT-135 spectra indicated the presence of thirty carbons atoms, consistent ]]> 30H26O17, the presence of seven methyl groups and eight carbonyl groups was also indicated. The complete structural elucidation of 7was achieved by analysis of the COSY and HMBC spectra. HMBC correlations were observed between d 2.57 (3H, s, H-8) and C-1 (d 119.6), C-5 (d 124.9) and C-6 (d 147.4); d 5.08 (1H, d, Ha-9) correlated with C-2 (d 145.6), C-3 (d 126.1), C-4 (d 148.9) and one carbonyl carbon; d 5.48 (1H, d, Hb-9) correlated with C-2 (d 145.6); d 7.99 (1H, s, H-9) correlated to C-2 (d 158.8), C-4 (d 152.5) and two carbonyl carbons; the aromatic proton d 6.97 (1H, s, H-5) correlated with C-8 (d 21.3), C-1 (d 119.6), C-3 (d 118.2) and C-4 (d 152.5), d 7.89 (1H, s, H-8) correlated with C-7 (d 163.6) and one carbonyl carbon. The methyl groups appear at 13C spectrum data at d 20.9, 20.8, 20.7, 20.7, 20.6 and 20.4, the carbonyl groups at d 170.5, 169.1, 169.0, 168.9, 168.3 and 168.1. In the 1H spectrum data at d 2.43, 2.35, 2.21, 2.14, 2.09 and 1.95 (Table 2). The structure was established as hexaacetylsalazinic acid, confirming that the compound salazinic acid was ]]> Everniastrum cirrhatum.
Table 2. 13C chemical shifts of decarboxythamnolic acid 3, thamnolic acid 4, baeomycesic acid 5, salazinic acid 6 and hexaacetylsalazinic acid 7
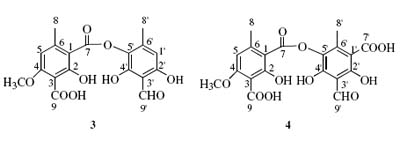
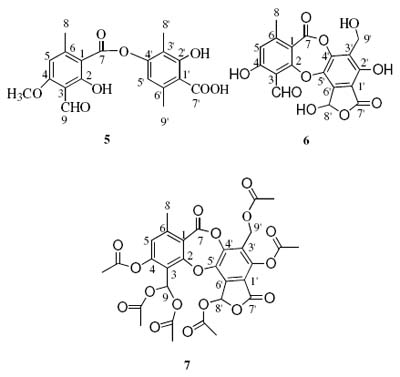
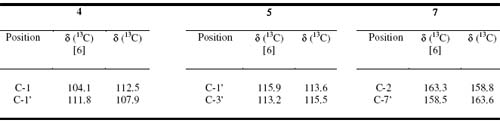
A comparison the assignment of the 13C signals of compounds 4, 5 and 7 was made with those reported in the literature6 and we found several differences (the chemical shifts of C-1 and C-1 in 4, C-1 and C-3 in 5, and C-2 and C-7 in 7). The correct assignment of the 13C shifts of the indicated carbons in compounds 4, 5 and 7 are therefore summarized in Table 3.
]]>Table 3. Comparison of 13C data of different chemical shift of 4, 5 and 7 with respect [6]
EXPERIMENTAL
General
The mass spectra (FAB) were recorded with a Jeol SX 102 spectrometer, 1H NMR (400 MHz) and 13C NMR (100 MHz), on a Bruker DRX 400 instrument. UV spectra were recorded with a Varian Cary 2290 specrometer.
Plant material
Everniastrum cirrhatum, Baeomyces roseus and Thamnolia vermicularis were collected in the Pongo region 3800 m.a.s.l. (La Paz Bolivia) in December 2002. The material was identified by department of Botany Herbario Nacional de Bolivia.
]]>Extraction and Isolation
Everniastrum cirrhatum was cleaned, dried at room temperature for 48 h and powdered. This material (156.0 g) was extracted with acetone (1 L) at room temperature for 72 h and concentrated in vacuo to yield the crude extrac (21.6 g) which was washed three times with CH2CL2 (500 mL) and EtOAc (500 mL). The soluble part was concentred under reduced pressure and subjected to chromatography. Atranorin (9.0 g) was obtained as the major product extract afforded a white solid 1 (9.0 g), which was recrystallized from ethyl acetate. From the CH2Cl2-EtOAc insoluble parts the depsidone 6 (8.4 g) was obtained.
Acetylation of 1. Compound 1 (400 mg) was dissolved in pure CHCl3 (8 mL). At room temperature and under stirring benzoyl chloride (902 mg) was dropped in followed by pyridine (2.5 mL). The reaction mixture was stirred for 3 h at room temperature and washed with water. Dried over Na2SO4 and evaporated. The residue was chromatographed with hexane/CHCl3 (1:0 to 0:1), 163 mg of 2 were obtained.
Acetylation of 6. Compound 6 (100 mg) in acetic anhydride (2.0 mL) and two drops of H2SO4 for 24 h at room temperature under stirring. After the usual work-up, 60 mg of 7 were obtained.
Baeomyces roseus was cleaned, dried at room temperature for 48 h and powdered. This material (36.0 g) was extracted with acetone at room temperature for 72 h and concentrated in vacuo to yield the crude extrac (2.6 g) which was recrystallized from acetone to gave a yellow solid 5.
Thamnolia vermicularis was cleaned, dried at room temperature for 48 h and powdered. This material (20.0 g) was extracted with acetone at room temperature for 120 h and concentrated in vacuo to yield the crude extrac (1.4 g) which was washed several times with Acetone each one (200 mL). This extract was subjected to CC on silica gel eluted with CH2Cl2-Acetone mixtures of increasing polarity. Two compounds were obtained, two meta-depsides 3 (89 mg) and 4 (1200 mg).
Atranorin (1) m.p. 197-198°C; FABMS m/z [M+1]+ 375 for C19O8H18; 1H NMR (CDCl3) and 13C NMR (CDCl3) see table 1.
2,4-bis(benzoyloxy)-atranorin (2); m.p. 171-172 °C; FABMS m/z [M+1]+ 583 for C33H26O10; 1H NMR (CDCl3) and 13C NMR (CDCl3) see table 1
Decarboxythamnolic acid (3), m.p. 216°C; FABMS m/z [M+1]+ 377 for C18H16O9; 1H NMR (DMSO-d6): d 2.50 (3H, s, H-8), 6.65 (1H, s, H-5), 2.28 (3H, s, H-8), 6.38 (1H, s, H-1), 10.23 (1H, s, H-9), 3.90 (3H, s, OMe-C-4); 13C NMR (DMSO-d6): (C-1) 112.4, (C-2) 166.9, (C-3) 105.8, (C-4) 161.5, (C-5) 106.3, (C-6) 145.1, (C-7) 161.0, (C-8) 22.3, (C-9) 170.5, (C-1) 108.2, (C-2) 159.9, (C-3) 109.8, (C-4) 154.0, (C-5) 129.4, (C-6) 143.4, (C-8) 17.9, (C-9) 194.9, (OCH3) 57.1.
]]>Thamnolic acid (4), m.p. 224°C, FABMS m/z [M+1]+ 421 for C19H16O11; 1H NMR (DMSO-d6): d 2.50 (3H, s, H-8), 6.13 (1H, s, H-5), 2.45 (3H, s, H-8), 10.4 (1H, s, H-9), 3.86 (3H, s, OMe-C-4); 13C NMR (DMSO-d6): (C-1) 112.5, (C-2) 166.8, (C-3) 105.5, (C-4) 161.7, (C-5) 106.5, (C-6) 145.4, (C-7) 164.5, (C-8) 22.3, (C-9) 170.5, (C-1) 107.9, (C-2) 161.0, (C-3) 109.5, (C-4) 157.3, (C-5) 129.9, (C-6) 143.9, (C-7) 172.5, (C-8) 16.4, (C-9) 194.9, (OCH3) 57.1.
Baeomycesic acid (5) m.p. 233 °C, FABMS m/z [M+1]+ 375 for C19H18O8; 1H NMR (DMSO-d6): d 2.47 (3H, s, H-8), 6.69 (1H, s, H-5), 2.51 (3H, s, H-8), 2.02 (3H, s, H-9), 6.45 (1H, s, H-5), 10.21 (1H, s, H-9), 3.96 (3H, s, OMe-C-4). 13C NMR (DMSO-d6): (C-1) 112.4, (C-2) 160.7, (C-3) 108.1, (C-4) 163.0, (C-5) 104.6, (C-6) 148.9, (C-7) 163.9, (C-8) 21.0, (C-9) 193.6, (C-1) 113.6, (C-2) 162.2, (C-3) 115.5, (C-4) 151.5, (C-5) 114.8, (C-6) 139.0, (C-7) 172.9, (C-8) 22.7, (C-9) 9.1, (OCH3) 56.1.
Salazinic acid (6) (11.1 g), m.p. 281-282 oC; FABMS m/z [M+1]+ 389 for C18H12O10; 1H NMR (DMSO-d6); d 2.51 (3H, s, Me-9), 4.58 (2H, s, CH2-9), 6.78 (1H, s, H-8), 6.81 (1H, s, H-5), 10.23 (1H, s, CHO-8). 13C NMR (DMSO-d6): d (1) 112.8, (2) 164.4, (3) 111.5, (4) 164.9, (5) 118.3, (6) 153.2, (7) 161.1, (8) 22.3, (9) 193.6, (1) 110.5, (2) 153.7, (3) 124.3, (4) 149.0, (5) 139.0, (6) 138.2, (7) 166.7, (8) 96.0, (9) 53.6.
Hexaacetilsalazinic acid (7); m.p. 374°C; FABMS m/z [M+Na]+ 681 [658+23] for C30H26O17; 1H NMR (CDCl3): d 2.55 (3H, s, H-8), 5.10 (1H, d, CH2-8), 5.4 (1H, d, CH2-8), 7.80 (1H, s, H-8), 6.90 (1H, s, H-5), 7.95.0 (1H, s, H-8). 13C NMR (CDCl3):d (1) 119.6, (2) 158.8, (3) 118.2, (4) 152.5, (5) 124.9, (6) 147.4, (7) 159.6, (8) 22.3, (9) 84.3, (1) 116.7, (2) 145.6, (3) 126.1, (4) 148.9, (5) 143.3, (6) 137.1, (7) 163.6, (8) 90.6, (9) 55.2.
Acknowledgements
]]> The authors wish to express their gratitude to the Swedish International Development Cooperation Agency (Sida) for financial support.
REFERENCES
1. Purvis, W. Lichens, Smithsonian Institution, Washington,D.C in association with The Natural History Museum, Press: London 2000. [ Links ]
2. Hale, M., “The Biology of lichens” Third Edition, Ed. Spottiswoode Ballantyne Ltd., Press: London 1983.
3. Dayan, F., Romangi, J., Pesticide Outlook 2001, 229-232. [ Links ]
4. Macia, M., Garcia, E., Vidaurre, P., Journal Ethnopharmacology 2005, 97, 337-350. [ Links ]
5. Sundholm, G., Huneck, S., Letter to Chemica Scripta 1980, 16, 197-200. [ Links ]
6. Sundholm, G., Huneck, S., Letter to Chemica Scripta 1981, 18, 233-236. [ Links ]
7. Eifler-Lima, V., Sperry, A., Sinbandhit, S., Boustie, J., Tomasi, S., Schenkel, E., Magnetic Resonance in Chemistry 2000, 38, 472-474. [ Links ] ]]>