DNA AFFINITY SCREENING OF PLANTS FROM THE BOLIVIAN AMAZONAS RAIN FOREST
Marcelo Bascopea b*, Jose Vila a c d e, Gilda Erosa a and Olov Sternera.
aDepartment of organic chemistry, Lund University, Lund, Sweden.
bCentro de Tecnología Agro industrial, Universidad Mayor de San Simón
cInstituto de Investigaciones Químicas (IIQ), Carrera de Ciencias Químicas, Universidad Mayor de San Andrés, (UMSA) Casilla 303, La Paz- Bolivia.
dInstituto de Investigaciones en Productos Naturales (IIPN, UMSA), Casilla 303, La Paz- Bolivia.
Keywords: Crude extracts, DNA affinity, antitumor, antimalarial, HPLC, Bolivian Amazonas.
ABSTRACT
Six crude extracts of plants from the Bolivian Amazonas rainforest were subject of DNA affinity screening, to identify potential antitumor and antimalarial activity, the assay uses a HPLC method, based on the ability of antitumor agents to reduce DNA peak size. Three extracts showed positive peak reduction, two were negative and resulted inconclusive, further observations indicate that the presence of the DNA in the mixture also reduces or eliminates substances peaks in the Chromatogram, possibly indicating specific affinity.
Seis extractos crudos de plantas de la amazonia Boliviana se sometieron a pruebas de afinidad de ADN, para identificar potencial actividad anti tumoral y anti malaria, el ensayo utiliza un método en HPLC, el cual se basa en la habilidad de los agentes anti tumorales para reducir el tamaño del pico de ADN. Tres extractos mostraron reducción positiva del pico, dos fueron negativos e inconclusos, Otras observaciones indicaron que la presencia del ADN en la mezcla también reduce o elimina los picos de las sustancias del extracto en el cromatograma, posiblemente indicando afinidad especifica
INTRODUCTION
One mechanism of action of different active substances involves interaction with DNA, several antitumor and antimalarial agents use this mechanism. Petuzzo et, al 1991 developed a rapid and efficient method for preliminary determination of DNA interaction for pure compounds and crude extracts, the method uses a HPLC as an affinity probe, using the ability of binding compounds to reduce the size of the DNA peak. Successful application of these method have been reported to identify active alkaloids from Arisarum vulgare by Melhaoui and Belouali in 1998 and in 1999 by Abourriche et. al. to investigate extracts from Cynthia savignyi a tunicate from the Moroccan atlantic sea. In our attempt to implement a fast and cheap preliminary assay to identify potential antitumor and antimalarial active plants from the bolivian amazonas rain forest, crude extracts from 6 plants were screened.
EXPERIMENTAL SECTION
Plant material and preparation of crude extracts.
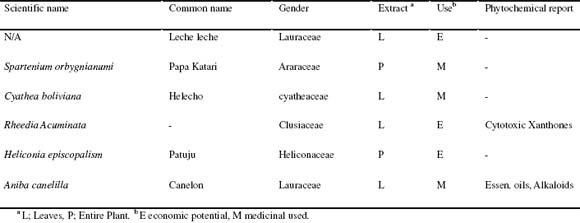
The six crude extracts used for the study, are part of an ongoing research of secondary metabolites of Bolivian plants, three of them have some reports concerning use in the traditional medicine, and the remaining three are considered to have some potencial economic use table 1 sumarizes the species, use and phytochemical reports concerning the plant. Samples were preferably from different families.
Plants were collected in the Valle de Sacta and the Siberia part of the Carrasco National park in Cochabamaba, Bolivia, during January 2003. The identification was carried out by the National herbarium Martin Cardenas Cochabamba, were a voucher of each specimen of the plant is deposited. Air dried powdered leaves, or entire plant were macerated with 96% ethanol three times during 24 Hrs each time at room temperature, after the solvent was removed under reduced pressure to obtain a crude extract.
Table 1. Plants screened and related reports.
]]>
DNA affinity test.
Calf Thymus DNA was purchased from Fluka, Ethidium Bromide was purchased from Acros organics. HPLC, The HPLC instrument used was a Dynamax Rainin model SD-200, equipped with an absorbance detector Rainin Model UV-1, and a Dynamax-100A, the column used was a Lichrospher RP-18 (5μm) 4X125 mm. DNA solution (0.1 mg/ml) were prepared in H2O and divided in sterilized vials containing 1 ml of solution which were stored at -20 C until use.

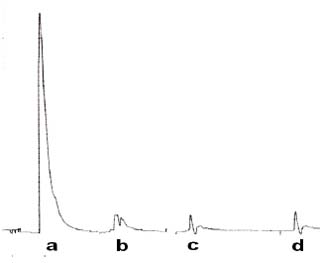
Figure 1. Comparative size of peaks for DNA affinity analysis; a) DNA alone, b) Solvent Alone (water or Methanol both have similar size), indicating negligible interference, c) co injection with ethidium Bromide 5 min incubation, d) 10 min incubation.
Crude extracts were dissolved in MeOH (1.0 mg/ml), solutions were centrifuged and filtrated to avoid injection of solids to the column. The mobile phase system used was H20 MeOH (80-20) during 5 minutes and gradient to pure MeOH in 5 minutes, rinse of the column for 20 minutes with MeOH before the next injection, equilibration time was 10 minutes, flow rate 1 ml/min and injection volume was 20 μl. Premixing of the sample with the DNA solution (v:v, 1:1), and incubation of the mixture at room temperature for 5-30 min. were carried on before injection. In these conditions free DNA eluted in approximately 1.8 min. In all experiments an injection of DNA alone coming from the same vial used to incubate the sample was made and the size of the eluted peak was used to compare with the one from the injection of the incubed mixture. All the samples were tested using the methodology described above, to increase the confiability of the procedure, crude extracts were tested three times.
To confirm the efficiency of the method used, ethidium bromide was used as positive control of the binding with DNA, almost total reduction of the DNA peak was obtained when co injected with the DNA solution as seen in figure 1.
RESULTS, DISCUSSION
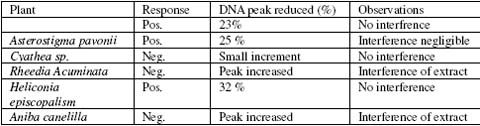
]]> Influence of solvents co injected (water, methanol and DMSO) in the DNA peak size profile, were studied by injecting pure solvents, and observing the retention time and the size of the peak produced by them in the chromatogram, water and Methanol elute at the same retention time of the free DNA, but the size of the peaks produced by them is less than the 5 % of the DNA peak size, discarding in this way interference by solvents, and the Void volume. DMSO shows a peak extremely big but the retention time is different with the free DNA, co injection of DNA and DMSO shows two peaks barely overlapped close to the base line, discarding in this way positive interference of the DMSO peak. No negative interference was observed. When no DNA reduction peak was observed, or an increment in the size of the DNA peak found, pure sample diluted with equal volume of water was injected to observe possible positive interference of sample compounds eluting at the same retention time of the free DNA. Positive control of the test with ethidium bromide shows and immediate reduction of the DNA peak after 5 minutes of incubation, The DNA peak is reduced to its 6 %, this signal could be considered as solvent signal or disruption of the baseline by the injection or void volume. Three of the extracts showed significant reduction of the DNA peak size, two extracts showed no reduction on the DNA peak and one of the test resulted as inconclusive.Table 2. Results of screening for DNA interaction of crude extracts.
With this results we can prioritize our studies on the plants which showed DNA affinity, and the method is fast, and it could allow to screen several samples in a short period of time, helping in this way to discrimine between potential plants with DNA affinity activity, even if the plant doesnt have reports as a medicinal plant.
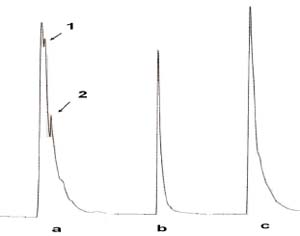
Figure 2. Elimination of peaks from crude extract by DNA affinity, a) Crude extract injected alone same retention time of b) free DNA resulting in positive interference of the c) coinjected sample +DNA. Note that peaks 1 and 2 are eliminated indicating possible specific DNA affinity with those compounds
An interesting behavior was observed when a sample of the gender Melastomataceae (plant not completely identified), was tested under the same conditions as the previous crude extracts. When the extract is injected alone, it presents three slightly overlapped peaks very close to the retention time of the free DNA see figure 2. But when co-injected with DNA, only one peak is visible, and this peak isnarrower than the crude extract peak alone, indicating in this way that, even if it is not possible to determine the DNA reduction due the interference from the crude extract. It would be possible to identify possible DNA binding analyzing retention times and size of the remaining peaks, based on the principle that a binded substance will behave in a total different way than the pure substance changing its retention time in the chromatogram.
ACKNOWLEDGEMENTS
]]> We want to thank Dr. Jorge Quillaguaman, Biotechnology department Lund University (Sweden) for DNA preparation. The authors are grateful to the Swedish International Development Agency (Sida) and its research division SAREC, for the economical support of this work.
REFERENCES
1. Melhaoui A, Belouali H (1998) DNA affinity of active alkaloids from Arisarum vulgare Targ. Journal of Ethnopharmacology 1998,62, 67-71
2. Abourriche A.; Charrouf M.; Bennamara A.; Berrada M.; Chaib N.; Boudouma M.; Francisco C.Journal of Ethnopharmacology, 1999. 68, 47-53 [ Links ]
3. Pezzuto J., Chun-Tao Che, McPherson D, Ji-Ping Zhu, Gulacti Topcu, Erdelmeier C., Cordell G., J. Nat. Prod., 1991, 54 (6), pp 15221530 [ Links ] ]]>