Ecología en Bolivia
versión impresa ISSN 1605-2528versión On-line ISSN 2075-5023
Ecología en Bolivia vol.52 no.2 La Paz set. 2017
Artículo
Associating immatures and adults of aquatic insects using DNA barcoding in high Andean streams
Asociando inmaduros y adultos de insectos acuáticos utilizando códigos de barras de ADN en ríos altoandinos
Carlos I. Molina1,2*, François-Marie Gibon3, Eduardo Dominguez4, Thomas Pape2 & Nina Rønsted2
1Instituto de Ecología, Universidad Mayor de San Andrés, Casilla 10077 - Correo Central, La Paz, Bolivia. Author correspondence: camoar6088@gmail.com
2Natural History Museum of Denmark, University of Copenhagen, Øster Voldgade 5-7,1350 Copenhagen, Denmark.
3Centre de Biologie pour la Gestion des Populations (UMR INRA / IRD / CIRAD / Montpellier Supagro), Campus de Baillarguet, CS 30016, 34988 Montferrier sur Lez Cedex, France.
4CONICET, Instituto de Biodiversidad Neotropical, Facultad de Ciencias Naturales IML, Universidad Nacional de Tucumán, San Miguel de Tucumán, Tucumán, Argentina.
Abstract
We analyzed the feasibility of using DNA-barcoding as a tool to achieve a correct and rapid association between different life stages of Ephemeroptera, Plecoptera, and Trichoptera insects in high Andean streams from La Paz, Bolivia. We focused on this particular environment because the water of streams from the glaciers is becoming scarce, and this could create a risk of local disappearance of these aquatic species. Using cytochrome c oxidase subunit 1 (COI) gene sequences, we found high genetic interspecific divergence between specimens from different families and genera (maximum 20%), whereas the intraspecific genetic divergences were lower between specimens of the same species (ranged from 0.1-2%). In this manner, we associate the larval or nymphal instar with their respective adults (female and male), for one species of mayfly (Meridialaris tintinnabula Pescador & Peters, 1987), three species of stoneflies (Anacroneuria vagante Stark & Baumann, 2011, Claudioperla tigrina Klapálek, 1904 and C. ruhieri Gibon & Molina, 2013) and one species of caddisfly (Anomalocosmoecus cf. illiesi Marlier, 1962). We concluded that COI Barcoding can be an effective tool for associating life stages and this tool could speed up the exploring the biodiversity patterns of aquatic insects in the High Andean region.
Key words: Biodiversity pattern, DNA-barcode, High Andean region, Larval/nymphal-adult association, Taxonomy.
Resumen
Analizamos la viabilidad del uso de códigos de barras de ADN como una herramienta para lograr una asociación rápida y correcta entre las diferentes etapas de vida en algunos insectos de los órdenes Ephemeroptera, Plecoptera y Trichoptera, correspondiente a cursos de aguas corrientes de la región Andina de La Paz, Bolivia. Nos enfocamos en estos ambientes, debido a que el agua de los arroyos de los glaciares empieza a escasear, y esto podría crear un riesgo en la desaparición local de estas especies acuáticas. Por medio del uso de las secuencias del gen citocromo c-oxidasa subunidad 1 (COI), encontramos alta divergencia interespecífica genética entre muestras de diferentes familias y géneros (máximo 20%), mientras que las divergencias genéticas intraespecíficas fueron menores entre ejemplares de la misma especie (osciló entre 0.1-2%). De esta manera asociamos el estadio larval o de ninfa con sus respectivos adultos (hembras y machos), para una especie de efemeróptero (Meridialaris tintinnabula Pescador & Peters 1987), tres especies de plecópteros (Anacroneuria vagante Stark & Baumann 2011, Claudioperla tigrina Klapálek 1904 y C. ruhieri Gibon & Molina 2013) y una especie de tricóptero (cf. Anomalocosmoecus illiesi Marlier, 1962). Concluimos que el código de barras de ADN podría ser una herramienta eficaz para asociar etapas de la vida y esto podría acelerar el estudio de los patrones de biodiversidad de insectos acuáticos de la región altoandina.
Palabras claves: Asociación de larvas / ninfa-adulto, Código de barras de ADN, Patrón de la biodiversidad, Región altoandina, Taxonomía.
Introduction
Traditionally, the composition of macroinvertebrate communities has been used for assessing the health status of freshwater environments. Within aquatic macroinvertebrate communities, insects belonging to the orders Ephemeroptera, Plecoptera, and Trichoptera (EPT) are the most used bioindicators due to high sensitivity to changing water conditions (Armitage et al. 1983, Doledec & Statzner 2008, Moya et al. 2011), which also means that climate change and habitat fragmentation poses a particularly serious threat to EPT taxa (Sandin et al. 2014). Unfortunately, in spite of their abundance in running waters of the Andean region, their diversity and fantastic knowledge remain poorly documented at the species level in the biodiversity era.
EPT insects spend most of their life cycle as larvae or nymphs in the aquatic environment (Molina & Puliafico 2016). Since the formal description of these species usually is based on the morphology of adult males, it is often complex or even impossible to achieve good taxonomic resolution at the species level for samples of larvae or nymphs without rearing these insects to adulthood. However, with development and accessibility of molecular phylogenetic technology, it is possible to also determine the species of immature stages (Zhou et al. 2009, Ruiter et al. 2013). Through concerted efforts of systematists and ecologists, the taxonomic knowledge of South American aquatic insects has improved recently (Fernández & Dominguez 2001, Dominguez & Fernández 2009), but many species still remain known only from their adult life stage, or they are not known at all.
Obtaining a good taxonomic resolution for immature stages of aquatic insects has never been an easy task. The conventional techniques of the immature-adult associations require many samples and include field or laboratory rearing. However, these procedures are complicated and time consuming and do not always provide reliable results due to the manipulation of the live specimens (Molina et al. 2008, Silva et al. 2012). Obtaining a good taxonomic resolution for immature stages will allow for more detailed ecological studies and permit a closer monitoring of species turnover caused by the subtle changes associated with climate changes.
Today, DNA barcoding is mostly providing a framework for clarifying the taxonomy of poorly known groups and used as an identification tool. For insects, good species delimitation can often be achieved by DNA barcoding using a short fragment of the mitochondrial cytochrome c oxidase subunit I (COI) (Hebert & Gregory 2005, Pons et al. 2006). Given an adequate database of reference sequences or DNA library, sequences can be used to group unidentified individuals (or even parts of these) with a priori-defined taxonomic entities based on similarity of the sequences.
The applicability of COI for species identification has been examined in several aquatic insect groups (Hogg et al. 2009, Zhou et al. 2009, Gattolliat & Monaghan 2010, Ruiter et al. 2013), but the shortage of sufficiently extensive databases of reference sequences hinders the possibility of associating sequences at lower taxonomic levels (Gattolliat & Monaghan 2010). In the Andean tropical region, DNA barcoding of aquatic insects has been scarcely attempted to date (Rozo-Lopez & Mengual 2015), but could potentially accelerate the knowledge of both adults and immatures, as vividly exemplified by Strutzenberger et al. (2011), who increased local species richness of geometrid moths in a well-known Andean mountain forest by 50%.
In this paper, we evaluate the feasibility of using DNA barcodes for the larval/nymphal-adult association for some EPT species of the high Andes. With the shrinking of the tropical glaciers leading to water shortage in this region (Soruco et al. 2009, Francou & Vincent 2010), we believe it is imperative and highly timely to develop the use of DNA barcoding of indicator species to more efficiently monitor this particular environment (Molina et al. 2008, Gibon & Molina 2013).
Methods
An aquatic insect survey was conducted over several days in 2014, sampling in different streams between the Apolobamba and Quimsa Cruz mountains (La Paz Department, Bolivia). Collected specimens were preserved in 95% ethanol. Larvae/ nymphs and adults were preliminarily sorted to family and genus using a stereo microscope and the morphological keys provided by Dominguez & Fernández (2009). We focused on obtaining reliable species identifications of the adult specimens. For this, one pair of legs, head, thorax, abdomen, wings and genitalia were dissected and mounted in Euparal on a slide. Another pair of legs was preserved in 95% of ethanol and kept at -18°C until DNA extraction. The identified, slide-mounted specimens are deposited in the Entomological collection of the Natural History Museum of Denmark, University of Copenhagen (ZMUC).
Molecular analysis
DNA was extracted using the DNeasy® Blood & Tissue kit (Qiagen, CA, USA) according the manufacturer's instructions and a 658 base pair (bp) long region of the cytochrome c oxidase subunit 1 (COI) gene was amplified and sequenced using the forward primer LCO1490 GGTCAACAAATCATAAAGATATTGG; and the reverse primer: HCO2198 TAAACTTCAGGGTGACCAAAAAATCA (Folmer et al. 1994). PCR amplification was performed as 25 µl reactions with 2 µl of DNA template, 2.5µl TQ, 10 µl dNTP mix, 2.5 µl each of primers LCO1490, and HCO2198, 0.1 µl DNA polymerase and 6 µl of ultrapure water. The PCR program consisted of an initial premelt of 94°C 1 min and 35 cycles following the profile: denaturing step at 95°C for 30 s, annealing step at 50°C for 30 s, extending step at 72°C for 54 s and finally the sample was stored at 4°C. PCR products were checked by Agarose gel electrophoresis (Invitrogen). DNA purification was performed with the Qiagen kit QIAquick® (Qiagen, CA, USA) following the manufacturer's protocol. Direct sequencing of purified PCR products was performed using BIGDYE v 1.1 (Applied Biosystems, Wellesley, Massachusetts, U.S.A.) and purified sequencing products were run on an AB3130x1 automated sequencer (Applied Biosystems).
Data analysis
The forward and reverse sequences were edited and assembled using Sequencher (2011). All the sequences were aligned in MEGA v 6.0.6 (Tamura et al. 2013), using the ClustalW algorithm with default parameters and according to the Drosophila yukaba sequence (Folmer et al. 1994, GenBank accession No. X03240). In order to evaluate the interspecific and intraspecific genetic divergences among our morphological species identified, we obtained other genetic sequences for a close related genus or same genus according to the pairwise alignment similarity (GenBank® database: www.ncbi.nlm.nih.gov/genbank/). All genetic sequences were subjected to pairwise nucleotide sequence divergence calculations using the Kimura 2-parameter (K2P) model. The species boundary associating were defined using the criteria that less than 2% genetic divergence indicates conspecificity (Hebert et al. 2003, Hogg et al. 2009). Finally, we built a gene tree using a Neighbour-joining (NJ) method with pairwise deletion of missing sites and K2P distance options (Kimura 1980).
All sequences were deposited under the project "Barcode Andean Aquatic Insects" (BAAI) in the Barcode of Life Data Systems (BOLD) (Ratnasinghan & Hebert (2007); www. barcodeoflife.org; sequence pages BAAI0002-15-BAAI0016-15) and cross-referenced to GenBank®.
Results
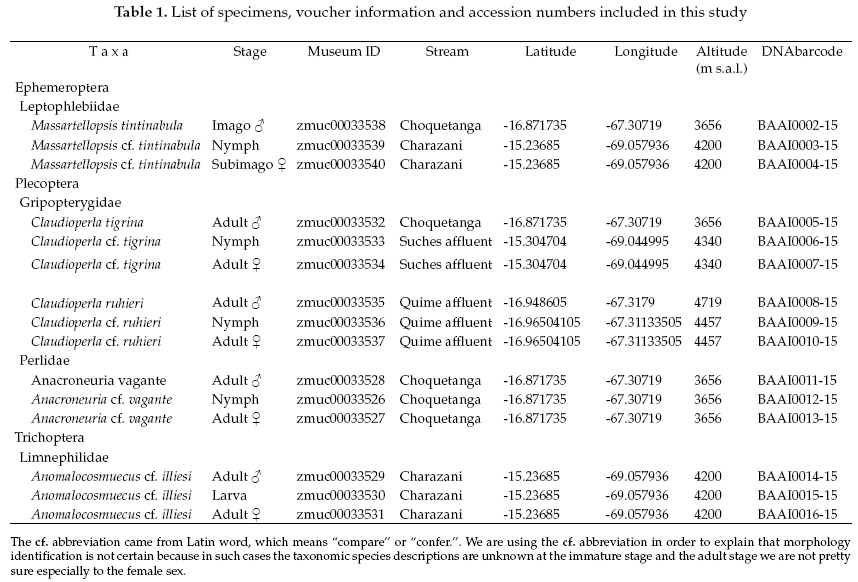
The geographic references of the specimens identified are shown in Table 1. DNA sequences from adult males identified to species based on morphology were used to explore their possible female and larval/nymphal complement specimens in order to verify the associations.
An average of 556 base pairs of the COI gene was recovered from 15 specimens. Compared with Drosophila yukaba sequences, our alignment covered a region between bp positions 1571 and 2127.
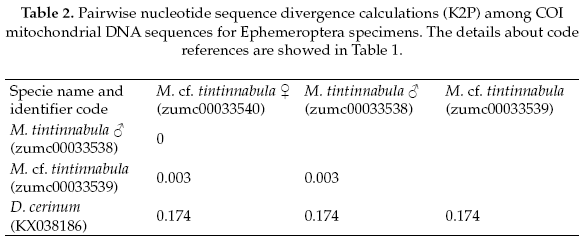
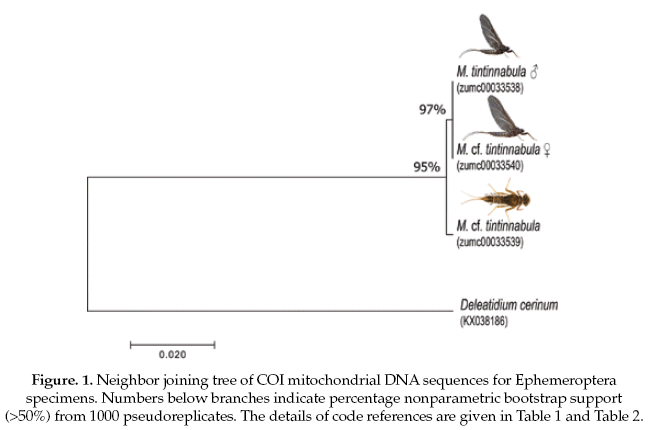
The Ephemeroptera specimens, the genetic sequence alignments we compared with a genetic sequence of the closely related genus: Delatidium cerinum (GenBank accession No.KX038186), which one showed 84% of pairwise identical with our alignments. The interspecific genetic divergences showed an average of 17% and the interspecific divergence was 0.2% (Table 2).
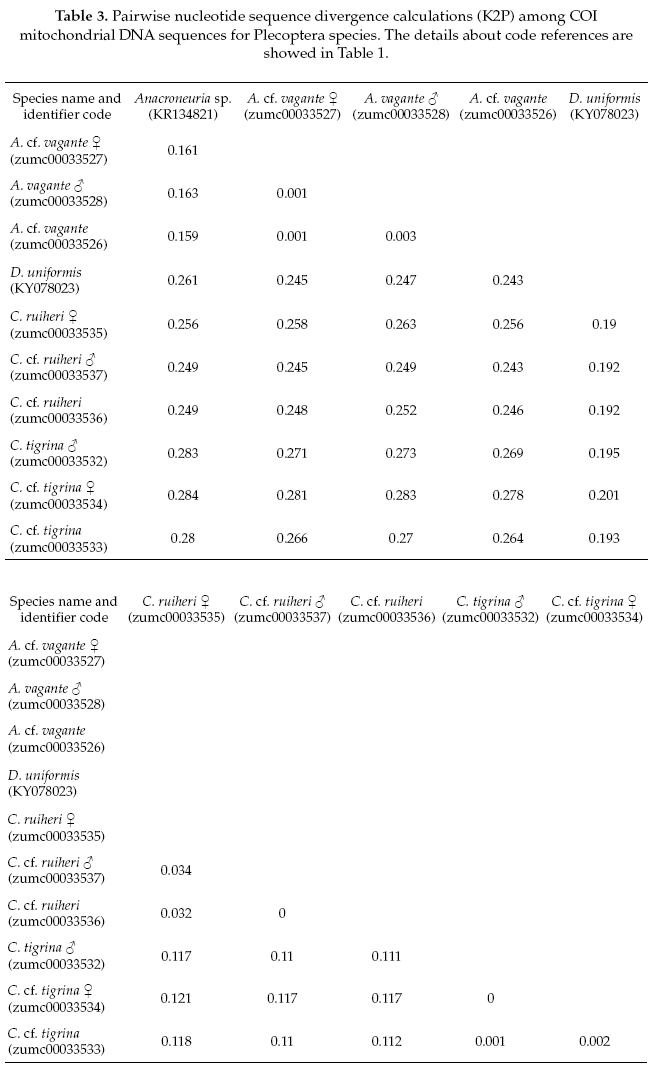
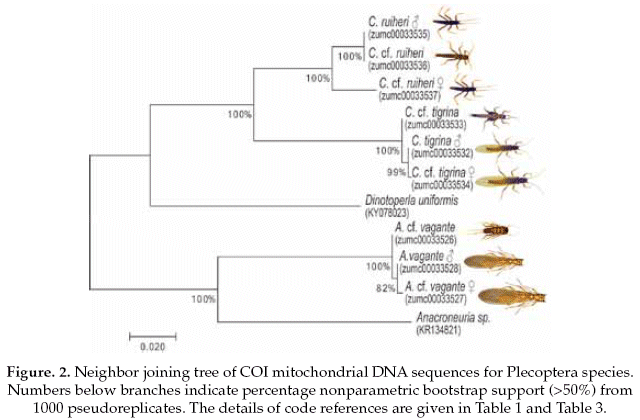
The Plecoptera species, on one side Anacroneuria vagante were compered with the same genus (Anacroneuria sp. GenBank accession No.KR134821), and showing 86% of pairwise alignment similarity. The interspecific genetic divergences were on average 16% and the interspecific divergences were also 0.2%. On the other side, the Claudioperla species were compared with close related genus (Dinotoperla uniformis, GenBank accession No.KX078023), with 83% of pairwise alignment similarity. The interspecific and intraspecific genetic divergences were 19% and 2%, respectively (Table 3).
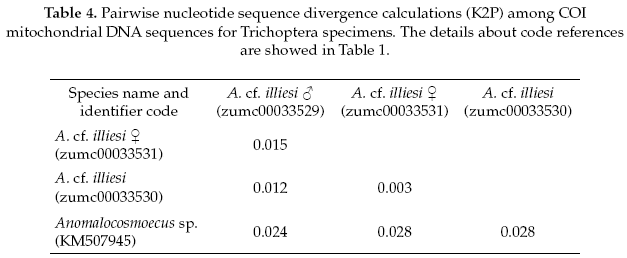
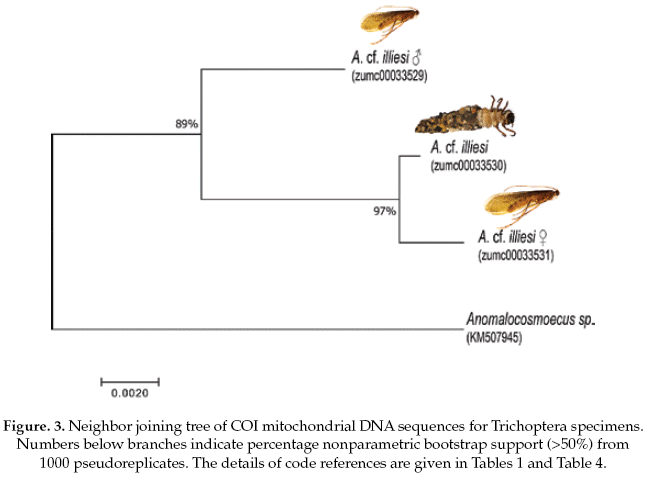
The Trichoptera specimens were compared with a specimen the same genus (Anomalocosmoecus sp., GenBank accession No.KM507945), showing 97% of pairwise alignment similarity and likely belong to close related species. The interspecific and intraspecific genetic divergences were 3% and 1%, respectively (Table 4).
In general, small intraspecific genetic divergences we observed among specimens of different developmental stages and among specimens of different sexes in the same species (genetic divergence range from 0.1% to 2%).
Based on the NJ analysis we were able to associate the larval/nymphal instars with their conspecific adults for one species of Ephemeroptera (Fig 1), three species of Plecoptera (Fig 2), and one species of Trichoptera (Fig 3). Three specimens (one larva/nymph, one adult male, one adult female) were always associated in a clade with high bootstrap support (100%). As these specimens show a <2% divergence, they are here considered as conspecific.
Discussion
To reliably identify specimens for precise species level delimitation, an extension of existing databases with genetic sequences (Gattolliat & Monaghan 2010) with data for EPT insects from the Andean region is needed. The two species of the genus Claudioperla Illies, 1963 (Plecoptera) showed an interspecific divergence <20%, and each had maximum intraspecific divergences <2% (Table 3). Claudioperla tigrina (Klapálek 1904) is a well-known species throughout the Andes, whereas the recently described C. ruhieri Gibon & Molina is known only from Bolivia (Gibon & Molina 2013).
The adult and immature Anacroneuria vagante Stark & Baumann, 2011 (Plecoptera) had a low interspecific divergence of 0.2%. This genus contains a large number of species, many of which were described from a single location or based on a single male specimen and without nymphal association. In Bolivia, 21 species of Anacroneuria are known (Stark & Baumann 2011), all without clear nymphal associations to the adult stoneflies.
Ephemeroptera and Trichoptera specimens showed low intraspecific divergence (1%). The Ephemeroptera species was difficult to identify even at genus level based on nymph and female subimago specimens. The single male imago that was obtained in the Choquetanga stream was identified as Meridialaris tintinnabula Pescador & Peters.
The classical taxonomic key for nymphs of this group indicate a strong similarity between this species and the single species of Massartellopis Demoulin, 1955 (Dominguez et al. 2006), and for this reason, identifications based only on nymphal and subimaginal instars are highly uncertain. The utility of morphology traits is clearly restricted with this example where nymphal characteristics are not diagnostic.
The Trichoptera specimens were readily identified to Anomalocosmoecus cf. illiesi Marlier based on the distinctive features given by Flint (1982). However, both male and female genitalia of this species have no known differences from the congeneric species A. blancasi Schmid, and these species could potentially be sympatric beyond their currently known co-occurrence in Lake Titicaca (Flint 1982). These difficulties suggest that more morphological and molecular studies are needed in order to clarify the systematics and distribution of these species.
PCR amplification success largely depends on choosing a suitable primer or set of primers. Previous studies of COI barcoding of EPT insects usually combined two primer sets (LCO1490/HCO2198 and LepF1/LepR1, Gill et al. 2014) or even included a third primer set for some caddisfly species (COI 2191/COI 1709, Zhou et al. 2009). In our study we found that the primer set of Folmer et al. (1994) (LCO1490/HCO2198) worked very well for all specimens analyzed and no differences were observed in sequence success for different life stages and sexes.
During our fieldwork, we obtained several immature aquatic insect but their adults were a weak collection due the hard climate condition of the high Andean mountain (strong wind and high humidity), because these adults are terrestrial flyers. The main reason for this is because we associated few specimens of the some exemplar EPT insects, but through of COI barcoding we demonstrate the reliability of associations with a specimen among larval/ nymphal and their respective adults at species level. We advocate establishing barcode sequence libraries for aquatic Andean insects to allow associations of specimens of different instars with their adult males and females. With eDNA surveillance (Thomsen et al. 2012, Bohmann et al. 2014) and next generation metabarcoding (Zhou et al. 2012) now being developed and refined, and with material costs for DNA barcodes rapidly decreasing (Meier et al. 2015), the major obstacle would seem to be funding and implementing the necessary large-scale inventories needed for securing a proper voucher material. Improved taxonomic knowledge of immature stages will lead to insights in their biology and allow for elaborating and testing of ecological hypotheses. Similarly, improving our knowledge and understanding of biodiversity patterns will enable us to detect and meet potential risks, e.g., as a result of climate change, in the vulnerable lotic ecosystems of the high Andean region.
Acknowledgments
Thanks Charlotte Hansen, David K.B. Cheung and Sree Gayathree from Natural History Museum of Denmark, for their assistance during the morphological diagnosis and molecular work of the aquatic insects. Thanks to staff at the IRD (French research institution in La Paz, Bolivia) for support to collect samples. Special thanks to Carla Maldonado, Natural History Museum of Denmark, for comments and improvements made to an earlier version. Thanks also to Kenneth Puliafico, USDA Forest Service for important suggestions and corrections to the manuscript.
References
Armitage, P. D., D. Moss, J. F. Wright & M. T. Furse. 1983. The performance of a new biological water quality score system based on macroinvertebrates over a wide range of unpolluted running water systems. Water Research 17: 333-347.
Bohmann, K., A. Evans, M. T. P. Gilbert, G. R. Carvalho, S. Creer, M. Knapp, D. W. Yu & M. de Bruyn. 2014. Environmental DNA for wildlife biology and biodiversity monitoring. Trends in Ecology & Evolution 29: 358-367.
Doledec, S. & B. Statzner. 2008. Invertebrate traits for the biomonitoring of large European rivers: An assessment of specific types of human impact. Freshwater Biology 53: 617-634.
Dominguez, E., C. Molineri, M. L. Pescador, M. D. Hubbard & C. Nieto. 2006. Ephemeroptera of South America. pp. 626. En: Adis, J., J. R. Arias, G. Rueda-Delgado & K. M. Wantzen (eds.) Aquatic Biodiversity in Latin America. Pensoft, Sofia.
Dominguez, E. & H. R. Fernández. 2009. Macroinvertebrados bentónicos sudamericanos. Sistemática y biología. Fundación Miguel Lillo, Tucumán. 656 pp.
Fernández, H. R. & E. Dominguez. 2001. Guía para la determinación de artrópodos bentónicos sudamericanos. Universdad de Tucumán, Tucumán. 282 pp.
Flint, O. S. 1982. Studies of Neotropical caddisflies, XXX: Larvae of the genera of South American Limnephilidae (Trichoptera). Smithsonian Contributions to Zoology 355:1-30.
Folmer, O., M. Black, W. Hoeh, R. Lutz & R. Vrijenhoek. 1994. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology 3(5): 294-299.
Francou, B. & L. Vincent. 2010. Les glaciers à l'épreuve du climat. IRD, Marseille. 16 p.
Gattolliat, J. L. & M. T. Monaghan. 2010. DNA-based association of adults and larvae in Baetidae (Ephemeroptera) with the description of a new genus Adnoptilum in Madagascar. Journal of the North American Benthological Society 29: 1042-1057.
Gibon, F.-M. & C. I. Molina. 2013. Contribution to the knowledge of the Andean Stonefly genus Claudioperla Illies, with description of new apterous and micropterous species (Plecoptera: Gripopterygidae). Zootaxa 42:170-177.
Gill, B. A., R. A. Harrington, B. C. Kondratieff, K. R. Zamudio, N. L. Poff & W. C. Funk. 2014. Morphological taxonomy, DNA barcoding, and species diversity in southern Rocky Mountain headwater streams. Freshwater Science 33(1): 288-301.
Hebert, P. D. N., Cywinska, A., Ball, S. L., & deWaard, J. R. 2003. Biological identifications through DNA barcodes. Proceedings of the Royal Society of London. Series B: Biological Sciences, 270(1512), 313-321. [ Links ]
Hebert, P. D. N. & T. R. Gregory. 2005. The promise of DNAbarcoding for taxonomy. Systematic Biology 54(4): 852-859.
Hogg, I. D., B. J. Smith, J. C. Bank, J. C. Denks, J. R. Dewaard & P. D. N. Hebert. 2009. Testing use of mitochondrial COI sequences for the identification and phylogenetic analysis of New Zealand caddisflies (Trichoptera). New Zealand Journal of Marine and Freshwater Research 43: 1137-1146.
Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution 16:111-120. [ Links ]
Meier, R., W. Wong, A. Srivathsan & M. Foo. 2015. $1 DNAbarcodes for reconstructing complex phenomes and finding rare species in specimen-rich samples. Cladistics (Early View) doi: 10.1111/ cla.12115.
Molina, C. I., Gibon F.-M, P. J. & R. C. 2008. Estructura de macroinvertebrados acuáticos en un río altoandino de la cordillera real, Bolivia: variación anual y longitudinal en relación a factores ambientales. Ecología Aplicada 7(1,2): 105-116.
Molina, C. I., & Puliafico, K. P. 2016. Life cycles of dominant mayflies (Ephemeroptera) on a torrent of the high Bolivian Andes. Revista de Biología Tropical 64(1): 275-287. [ Links ]
Moya, N., R. M. Hughes, E. Domínguez, F.-M. Gibon, E. Goitia & T. Oberdorff. 2011. Macroinvertebrate-based multimetric predictive models for evaluating the human impact on biotic condition of Bolivian streams. Ecological Indicators 11: 840-847.
Pons,J., T. Barraclough, J. Gomez-Zurita, A. D. Cardoso, D.P., S. Hanzell, S. Kamoun, W. Sumlin & A. P. Vogler. 2006. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Systematic Biology 55(4): 595-609.
Ratnasinghan, S. & P. D. N. Hebert. 2007. BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Molecular Ecology 7: 355-364.
Rozo-Lopez, P. & X. Mengual. 2015. Mosquito species (Diptera, Culicidae) in three ecosystems from the Colombian Andes: identification through DNA barcoding and adult morphology. ZooKeys 513: 39-64.
Ruiter, D. R., E. E. Boyle & X. Zhou. 2013. DNA barcoding facilitates associations and diagnoses for Trichoptera larvae of the Churchill (Manitoba, Canada) area. BMC Ecology 13:1-39.
Sandin, L., A. Schmidt-Kloiber, J.-C. Svenning, E. Jeppesen & N. Friberg. 2014. A trait-based approach to assess climate change sensitivity of freshwater invertebrates across Swedish ecoregions. Current Zoology 60: 221-232.
Sequencher. 2011. The complete software solution for sequencing DNA. Page v5.0.1. Gene Codes Corporation. [ Links ]
Silva, F. L., S. Wiedenbrug, S. Trivinho-Strixino, C. S. Neubernde Oliveira & M. Pepinelli. 2012. Two new species of Hudsonimyia Roback, 1979 (Diptera: Chironomidae: Tanypodinae) from Neotropical region unveiled by morphology and DNA barcoding. Journal of Natural History 46:1615-1638.
Soruco, A., C. Vincent, B. Francou & G.J.F. 2009. Glacier decline between 1963 and 2006 in the Cordillera Real, Bolivia. Geophysical Research Letters 36:1-6.
Stark, B.P. & R.W. Baumann. 2011. Records of Anacroneuria (Plecoptera: Perlidae) from Bolivia and Paraguay with descriptions of three new species. Illiesia 7:182-191.
Strutzenberger, P., G. Brehm & K. Fiedler. 2011. DNA barcoding-based species delimitation increases species count of Eois (Geometridae) moths in a well-studied tropical mountain forest by up to 50%. Insect Science 18: 349-362.
Tamura, K., G. Stecher, D. Peterson, A. Filipski & S. Kumar. 2013. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Molecular Biology and Evolution 30: 2725-2729.
Thomsen, P. F., J. Kielgast, L. L. Iversen, C. Wiuf, M. Rasmussen, M. T. Gilbert, L. Orlando & E. Willerslev. 2012. Monitoring endangered freshwater biodiversity using environmental DNA. Molecular Ecology 21: 2565-2573.
Zhou, X., S. J. Adamowicz, L. M. Jacobus, E. R. DeWalt & P. D. N. Hebert. 2009. Towards a comprehensive barcode library for arctic life - Ephemeroptera, Plecoptera, and Trichoptera of Churchill, Manitoba, Canada. Frontiers in Zoology 6: 30 (1-9)
Zhou, X., Y. Li, S. Liu, X. Su, L. Zhou, M. Tang, R. Fu, J.Li & Q. Huang. 2012. Ultra-deep sequencing enables high-fidelity recovery of biodiversity for bulk arthropod samples without PCR amplification. GigaScience 2: 4.
Artículo recibido el: 27 abril 2017
Aceptado en: 17 julio 2017
Manejado por: Diana Silva.











 uBio
uBio